Dask in ztfimg#
The `dask <https://docs.dask.org/en/stable/>`__ library enables to distribute computing between cluster (or to multiprocess from your machine). This library is natively incorporated within ztfimg to enable you to work with thousands of images fast and without exploding your memory.
The most natural way to use dask is by setting the use_dask option to True while loading a datafile. data will be stored as a dask.array in place of a numpy.array.
.compute() and .persist() methods enable you to compute or persist the “dasked” objects.
Dask Cluster#
let’s create a dask cluster. For this example, we will create one from the local machine, but the core interest of dask is that all computing will automatically scale whatever cluster you have access to ; see dask jobqueue
[2]:
client.close()
[3]:
import dask
from dask.distributed import Client
client = Client()
with that, you are all set ! Open you localhost:8787
Dasking a science image#
Following the quickstart tutorial:
[4]:
import ztfimg
img = ztfimg.ScienceQuadrant.from_filename("ztf_20200924431759_000655_zr_c13_o_q3_sciimg.fits",
as_path=False, use_dask=True)
With dask, no actual computation is made while loading the data
The data attribute is a dask.array.
[5]:
img.data
[5]:
|
||||||||||||||||
and if you set as_path=False the ztfquery.get_file() function itself is delayed, so img.filepath is a delayed object. But filename is the filename you provided if loaded using self.from_filename.
[6]:
img.filename
[6]:
'ztf_20200924431759_000655_zr_c13_o_q3_sciimg.fits'
[7]:
img.filepath
[7]:
Delayed('get_file-c8bd23ce-446f-4fbe-ad1c-8239423db0b9')
remark that the meta data are still known because they are based on the input filename.
Data access#
All computations are delaued, so whatever comes out of get_data() will also be a dask.array
[8]:
data_clean = img.get_data(apply_mask=True)
data_clean
[8]:
|
||||||||||||||||
[9]:
data_clean.visualize()
[9]:
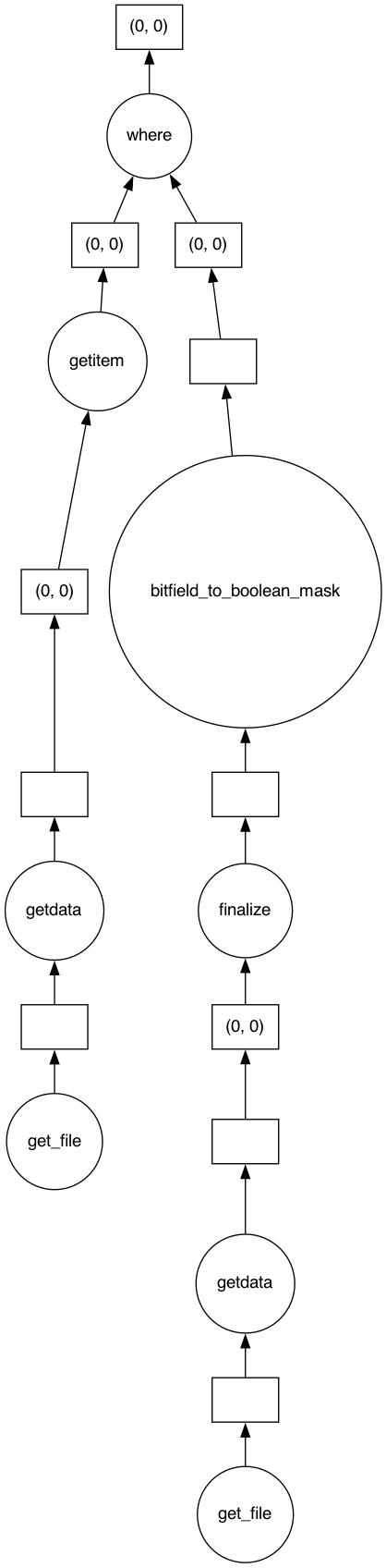
to get the actual array, simply do: .compute()
[10]:
data = data_clean.compute() # check you dask dashboard at localhost:8787
[11]:
data
[11]:
array([[ 44.91321 , 167.88715 , 206.81393 , ..., 185.91528 , 196.35838 ,
194.94452 ],
[ 16.670918, 78.78005 , 138.10672 , ..., 152.68452 , 149.92209 ,
158.07927 ],
[ 17.609827, 87.123825, 152.8368 , ..., 148.68439 , 154.78484 ,
167.03732 ],
...,
[ 16.900919, 76.01857 , 139.25365 , ..., 165.22946 , 159.50085 ,
154.41986 ],
[ 10.205329, 83.728935, 134.86244 , ..., 161.0656 , 147.19606 ,
155.4496 ],
[ 12.984478, 83.76391 , 133.77336 , ..., 160.41643 , 157.65231 ,
156.70863 ]], dtype=float32)
get_data().compute() vs. .compute()#
In the example above, the results of a function is computed but img remained “dasked”. To compute (in place) all dasked images attribute, use the compute() instance method.
[12]:
img.data
[12]:
|
||||||||||||||||
[13]:
img.filename
[13]:
'ztf_20200924431759_000655_zr_c13_o_q3_sciimg.fits'
[14]:
img.compute()
[15]:
img.data
[15]:
array([[156.70863 , 157.65231 , 160.41643 , ..., 133.77336 , 83.76391 ,
12.984478],
[155.4496 , 147.19606 , 161.0656 , ..., 134.86244 , 83.728935,
10.205329],
[154.41986 , 159.50085 , 165.22946 , ..., 139.25365 , 76.01857 ,
16.900919],
...,
[167.03732 , 154.78484 , 148.68439 , ..., 152.8368 , 87.123825,
17.609827],
[158.07927 , 149.92209 , 152.68452 , ..., 138.10672 , 78.78005 ,
16.670918],
[194.94452 , 196.35838 , 185.91528 , ..., 206.81393 , 167.88715 ,
44.91321 ]], dtype=float32)
[16]:
img.filepath
[16]:
'/Users/rigault/data/ztf/sci/2020/0924/431759/ztf_20200924431759_000655_zr_c13_o_q3_sciimg.fits'
[17]:
img.use_dask
[17]:
False
Stack bias images#
dask computing is particularly useful when dealing with many images. Let’s use here an example where we want to stack the 10 bias images of a given day.
[18]:
filename_biases = [
'ztf_20200801078252_000000_bi_c13_b.fits.fz',
'ztf_20200801078484_000000_bi_c13_b.fits.fz',
'ztf_20200801078727_000000_bi_c13_b.fits.fz',
'ztf_20200801078958_000000_bi_c13_b.fits.fz',
'ztf_20200801079201_000000_bi_c13_b.fits.fz',
'ztf_20200801079444_000000_bi_c13_b.fits.fz',
'ztf_20200801079676_000000_bi_c13_b.fits.fz',
'ztf_20200801079919_000000_bi_c13_b.fits.fz',
'ztf_20200801080162_000000_bi_c13_b.fits.fz',
'ztf_20200801080394_000000_bi_c13_b.fits.fz']
len(filename_biases)
[18]:
10
Opening one image without dask#
[19]:
bias = ztfimg.RawCCD.from_filename(filename_biases[0], as_path=False)
[20]:
_ = bias.show()
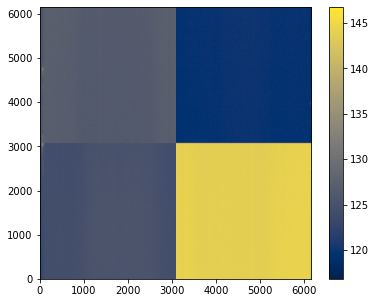

[21]:
data = bias.get_data(corr_nl=True, corr_overscan=True)
data
[21]:
array([[-0.3898886 , -3.0719259 , 1.1290246 , ..., -0.4186407 ,
0.3849024 , 1.4197382 ],
[-1.8835076 , -1.2695092 , 0.59942544, ..., 1.0178564 ,
1.09708 , -3.910992 ],
[-0.4443034 , 0.49627125, 0.3480245 , ..., -3.5802078 ,
-0.25536343, -1.1749039 ],
...,
[-1.7054825 , -2.6555634 , -2.0730667 , ..., 0.14506413,
-2.199937 , -1.9596875 ],
[-1.0927409 , -3.0640085 , 1.2626897 , ..., -0.30722025,
-3.4315488 , -2.5934827 ],
[-0.8860508 , -1.0225178 , 1.580769 , ..., -2.3171294 ,
0.13868105, -0.57472074]], dtype=float32)
using dask now#
[22]:
%time bias = ztfimg.RawCCD.from_filename(filename_biases[0], as_path=False, use_dask=True)
CPU times: user 7.34 ms, sys: 4.12 ms, total: 11.5 ms
Wall time: 10.7 ms
[23]:
bias.use_dask
[23]:
True
[24]:
bias.get_data()
[24]:
|
||||||||||||||||
As for quadrants, compute or persist the object
[25]:
bias.compute()
/Users/rigault/miniforge3/lib/python3.9/site-packages/ztfimg-0.15.10-py3.9.egg/ztfimg/base.py:859: UserWarning: compute of a collection is not optimal yet. Loops over images to call their compute.
warnings.warn("compute of a collection is not optimal yet. Loops over images to call their compute.")
[26]:
bias.use_dask
[26]:
False
[27]:
bias.get_data()
[27]:
array([[123.1784 , 120.49583 , 124.697624, ..., 142.85115 , 143.6549 ,
144.69003 ],
[121.684525, 122.298645, 124.16795 , ..., 144.28812 , 144.36736 ,
139.35794 ],
[123.12406 , 124.06482 , 123.91654 , ..., 139.68887 , 143.01462 ,
142.09483 ],
...,
[124.695724, 123.74546 , 124.32808 , ..., 118.84962 , 116.50417 ,
116.74445 ],
[125.30851 , 123.33685 , 127.66442 , ..., 118.397255, 115.27232 ,
116.11055 ],
[125.515175, 125.37868 , 127.98249 , ..., 116.38696 , 118.84325 ,
118.12971 ]], dtype=float32)
[28]:
bias.filenames
[28]:
['ztf_20200801078252_000000_bi_c13_b.fits.fz',
'ztf_20200801078252_000000_bi_c13_b.fits.fz',
'ztf_20200801078252_000000_bi_c13_b.fits.fz',
'ztf_20200801078252_000000_bi_c13_b.fits.fz']
[29]:
bias.filepaths
[29]:
['/Users/rigault/data/ztf/raw/2020/0801/078252/ztf_20200801078252_000000_bi_c13_b.fits.fz',
'/Users/rigault/data/ztf/raw/2020/0801/078252/ztf_20200801078252_000000_bi_c13_b.fits.fz',
'/Users/rigault/data/ztf/raw/2020/0801/078252/ztf_20200801078252_000000_bi_c13_b.fits.fz',
'/Users/rigault/data/ztf/raw/2020/0801/078252/ztf_20200801078252_000000_bi_c13_b.fits.fz']
Opening with dask and stacking them#
[30]:
%%time
biases = [ztfimg.RawCCD.from_filename(filename_, as_path=False, use_dask=True)
for filename_ in filename_biases]
CPU times: user 33.5 ms, sys: 3.85 ms, total: 37.4 ms
Wall time: 36.7 ms
all data loading (downloading) are delayed
[31]:
biases
[31]:
[<ztfimg.raw.RawCCD at 0x2a2626160>,
<ztfimg.raw.RawCCD at 0x2a2d06d30>,
<ztfimg.raw.RawCCD at 0x2a2d46280>,
<ztfimg.raw.RawCCD at 0x2a2d527f0>,
<ztfimg.raw.RawCCD at 0x2a2d5d370>,
<ztfimg.raw.RawCCD at 0x2a2d6f220>,
<ztfimg.raw.RawCCD at 0x2a2d78fd0>,
<ztfimg.raw.RawCCD at 0x2a2d8af40>,
<ztfimg.raw.RawCCD at 0x2a2d9ddf0>,
<ztfimg.raw.RawCCD at 0x2a2daecd0>]
[37]:
clean_data0 = biases[0].get_data(corr_nl=True, corr_overscan=True)
[38]:
clean_data0.visualize()
[38]:

[39]:
clean_data0.compute()
[39]:
array([[-0.38988861, -3.07192596, 1.12902466, ..., -0.41864069,
0.3849024 , 1.41973822],
[-1.8835076 , -1.26950918, 0.59942545, ..., 1.01785632,
1.09707995, -3.91099195],
[-0.44430339, 0.49627125, 0.34802449, ..., -3.58020779,
-0.25536343, -1.17490383],
...,
[-1.70548249, -2.65556336, -2.07306671, ..., 0.14506414,
-2.19993708, -1.95968745],
[-1.09274085, -3.06400855, 1.26268975, ..., -0.30722026,
-3.43154888, -2.59348278],
[-0.88605083, -1.02251781, 1.58076893, ..., -2.31712949,
0.13868106, -0.57472074]])
So to stack them, simply get all the dask arrays and sum/average them
[40]:
import dask.array as da
[41]:
%%time
stacked = da.stack([bias_.get_data(corr_nl=True, corr_overscan=True) for bias_ in biases])
# no computing yet
CPU times: user 232 ms, sys: 13 ms, total: 245 ms
Wall time: 245 ms
[42]:
averaged_data = da.mean(stacked, axis=0)
averaged_data
[42]:
|
||||||||||||||||
[43]:
averaged_data.visualize() # these are all the tasks to do
[43]:
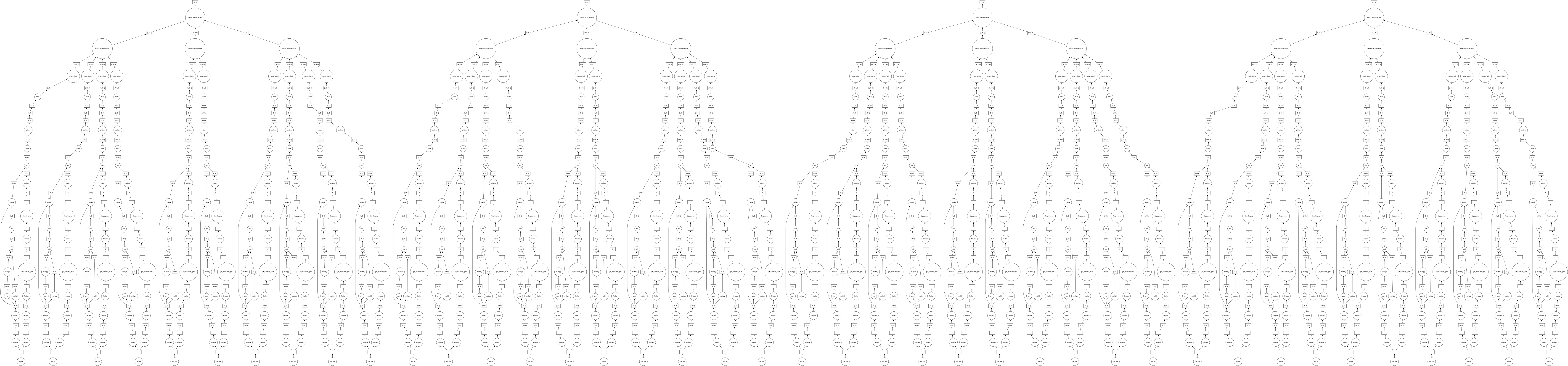
on now let’s run the computation. Dask will exploite the parallelism on the process (check the dask dashboard on localhost:8787)
[44]:
%%time
data = averaged_data.compute()
CPU times: user 684 ms, sys: 238 ms, total: 922 ms
Wall time: 4.28 s
[45]:
data
[45]:
array([[-1.1285973 , -1.717946 , -0.3112228 , ..., -0.7957014 ,
-0.96444684, -0.4791685 ],
[-1.4917496 , -0.11884248, -0.17186753, ..., -0.780125 ,
-0.29935104, -1.2285687 ],
[-0.2066387 , -0.98148984, -0.523502 , ..., -1.186869 ,
-1.0701668 , -1.1147332 ],
...,
[-0.06632094, -0.6073396 , -1.5902216 , ..., 0.05830754,
-0.9022645 , -0.5186822 ],
[-1.2467864 , -1.141419 , -1.2951422 , ..., -1.3223233 ,
-2.2815497 , -1.847585 ],
[-1.0298344 , -1.0996517 , -0.82145655, ..., -0.6130624 ,
-0.2943551 , -1.3127122 ]], dtype=float32)
careful if you are running this on your own machine (and not a cluster) the RAM will be limited to you single computer RAM, so do not open in // too many files.